This page contains an overview of NC State’s efforts and of PreMiEr’s Social and Ethical Implications (SEI) research focus. Visit the full PreMiEr website to learn more.
About
PreMiEr will develop an integrated framework that advances microbiome technologies and enables the bioinformed design of smart and healthy built environments.
The Center for Precision Microbiome Engineering (PreMiEr) is a National Science Foundation (NSF)-funded Engineering Research Center (ERC) (Award # 2133504) led by Duke University in collaboration with NC State University, North Carolina Agricultural and Technical State University (NC A&T), the University of North Carolina at Chapel Hill (UNC-CH), and the University of North Carolina at Charlotte (UNC-Charlotte), as well as members of industry and other educational institutions.
PreMiEr is funded by a five-year, $26 million grant, renewable for a second five-year, $26 million term.
Vision
PreMiEr’s vision is to develop an integrated framework for enabling the development of high impact microbiome technologies that provide innovative solutions to key societal challenges at the interface of human health and the built environment. In particular, PreMiEr will advance microbiome technologies by developing diagnostic tools and engineering approaches that lead to the prevention of infectious agents’ colonization and the promotion of beneficial microorganisms in the built environment.
Scope
Researchers in PreMiEr will achieve their goals through cross-disciplinary efforts using the latest technologies in genomic, transcriptomic, and metabolomic technologies to study the microbial “dark matter” that colonizes the built environment, develop sensors and other technologies to monitor and modify those communities, and create sophisticated computer models to help predict the outcomes of changes to the built environment microbiome and drive beneficial health outcomes.
Impact
PreMiEr seeks to help identify not only what might make a built environment microbiome harmful to inhabitants, but also hopes to identify organisms, metabolites, or other factors that lead to positive health outcomes. Our ultimate goal is to create biologically safe indoor spaces for everyone. The findings of this center could ultimately lead to recommendations for building design, construction, or operation in order to promote the proliferation of healthy microorganisms in man-made structures.
MoBE Workshop
MoBE – Workshop on the Societal and Ethical Implications of Microbiome Engineering in Built Environments
May 15 @ 9:00 am – 5:00 pm
Talley Student Union, Room 3222
This workshop is hosted by the GES Center at NC State and funded by the NSF Precision Microbiome Engineering (PreMiEr) research grant.
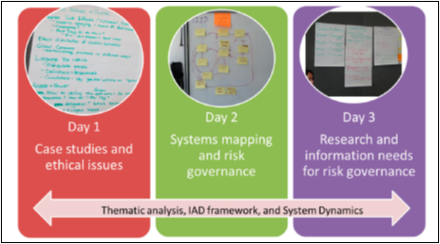
Wednesday, May 15: Speakers to discuss the State of MoBE Science and Engineering; Social Equity; Bioethics; Risk Governance; Public Perception and Community Engagement; and Integration of SEI with Science and Engineering of the Microbiome | Thursday, May 16: Presentation of case studies and deliberative discussions
Video
Watch our video, produced by Duke Engineering and originally posted on YouTube, to learn more >
Research Areas
Research Thrust 1 (RT1) combines multi-omic investigations to determine the mechanisms of microbial colonization and aims to develop sensor and tracking technologies for diagnosing built environment health at varying resolutions (i.e., personal, room, and building level).
RT1 researchers will develop tracking tools that combine phylogenetic and functional aspects through the integration of personal and environmental microbiome data with microbial dark matter characteristics. These tools will include genome-enabled approaches that can target uncultivated microbial taxa and increase our understanding of microbial diversity, phylogenetic relationships, metabolic capabilities, and interactions in the built environment as well as functional approaches via meta-omics. In combination, these approaches will deepen our databases enabling the identification of key molecules and the development of sensors for health assessment of the built environment.
Projects in RT1 will apply and expand fundamental knowledge in microbiome monitoring. We will begin by developing approaches for monitoring and connecting the personal and the environmental microbiome as well as determine functional signatures that can diagnose built environment health. RT1 data will provide the early building blocks for monitoring the built environment microbiome as well that of its occupants, identifying the biomarkers that signal a healthy built environment, and inform PreMiEr’s future sensor development work.
Projects in Research Thrust 2 (RT2) will build the toolbox needed for targeting the delivery/removal of desired genetic features or vectors in the built environment as well as enabling functional modulation in an established microbial community.
RT2 researchers will use their knowledge of delivery systems and nanoparticle transport in complex environments to develop the requisite toolbox for microbiome engineering in the built environment. Initial projects will focus on limiting the spread of animicrobial resistance (AMR) in built microbiomes as well as targeting the built environment water microbiome as there is a critical body of work linking the microbiome of premise plumbing to adverse effects in the built environment. Later efforts will transition to engineering solutions to limiting pathogens and bioaerosols as PreMiEr’s research matures.
Projects in Research Thrust 3 (RT3) will develop predictive models that incorporate spatiotemporal methods, generative modeling concepts, and machine learning approaches to analyze built environment microbiomes.
RT3 projects will focus on the development of predictive models that identify factors that contribute to microbiome compositional variations, and microbiome signatures that associate with specific health outcomes, which in turn will inform built environment health signature identification.
Initial projects will focus on various facets of the predictive models using existing large datasets and then incorporate PreMiEr datasets from Research Thrust 1 as those are generated. Spatial-temporal statistical models for microbiome compositions will be constructed that can characterize the personalized equilibrium of an individual’s microbiome compositions and detect anomalies, or deviation from the normal equilibrium. This information will be crucial in linking to the environmental microbiome. Another project will integrate functional information to decipher the functional meanings of the signals identified in the predictive models. Finally, improvements in the bioinformatic preprocessing pipelines using machine learning approaches will enhance the sensitivity and specificity of the predictive models.
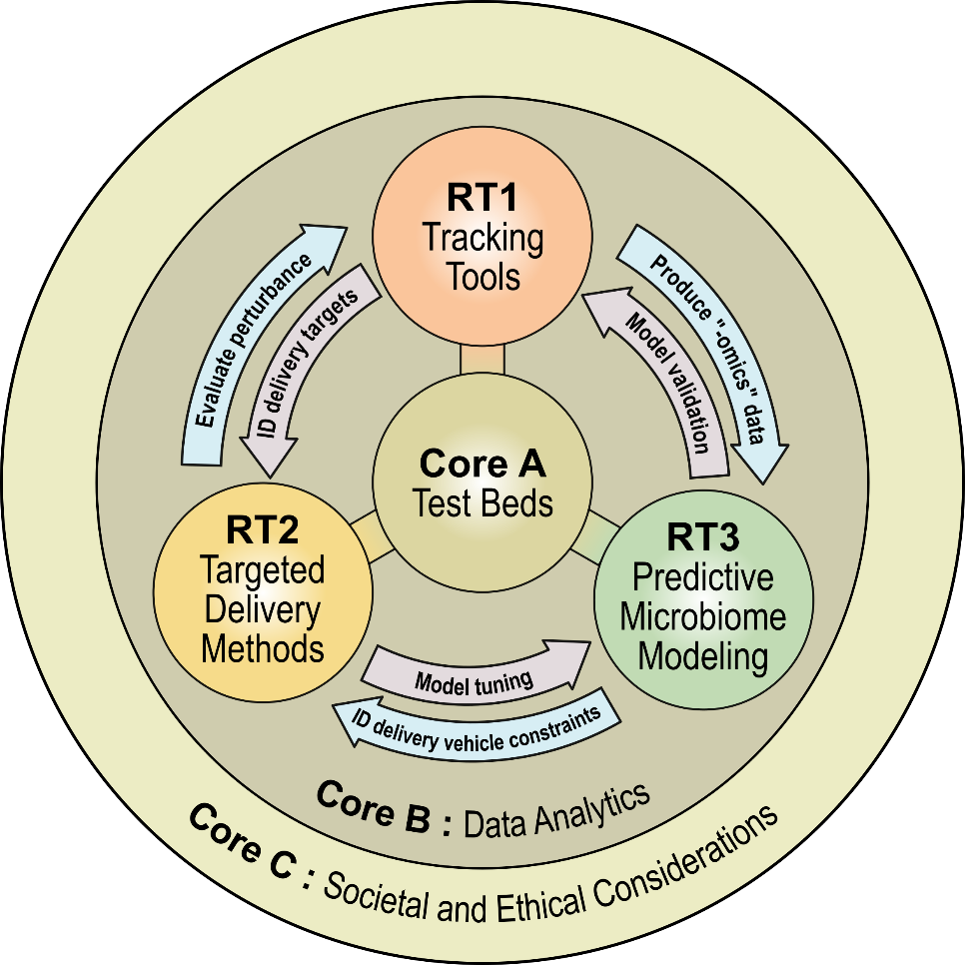
In Research Core A, PreMiEr’s microbiome tracking devices (personal and environmental sensors), targeted delivery tools and predictive microbiome modeling framework will be integrated to measure, predict and improve the health of the built environment microbiome in six model testbed environments:
- Environmental Chambers
- Artificial Gut
- Tiny House
- Duke University Smart Home
- Hospitals, and
- Bolivian Homes
The PreMiEr Data Analytics Core is an integral part of the success of the ERC by supporting the hardware and software needs of all other research thrusts. Its goals can be broken down into three main areas.
1. A seamless, central repository with core services providing
- Scientific data processing and analysis support
- Report and figure creation services
- Seamless and automated ability to start and modify processes and analyses
2. Ensuring transparency and reproducibility
- Eliminate “silos” of data
- Allow teams to easily work together
- Publish results that fully expose the processing pipeline and publish virtualized containers
- Support version control
3. Being “hardware agnostic”
- Allow any PreMiEr researcher (or other interested scientist) to faithfully reproduce analyses regardless of hardware
- Provide virtualized containers for on-campus and other systems, including a primary system at UNC-Charlotte, the ViCAR system at North Carolina A&T, and the cloud
Diversity and Culture of Inclusion Overview
PreMiEr prioritizes diversity and culture of inclusion (DCI) recognizing that our ERC cannot maximize impacts without intentional efforts to foster both and promote an equitable environment where all can thrive. PreMiEr grew out of an existing 5-year partnership between Duke (HWI) and North Carolina A&T (HBCU). This partnership taught both universities lessons on the practical implementation of diversity and culture of inclusion. The addition of North Carolina State (NC State), UNC Chapel Hill (UNC-CH), and UNC Charlotte (UNC-C) to this team has provided us with an even greater opportunity to institutionalize DCI in every aspect of PreMiEr’s function.
PreMiEr is dedicated to building a diverse faculty and trainee population as a means to broaden participation in STEM. This will be achieved by leveraging recruitment of underserved populations across all campuses and working to develop research pipelines between Duke, UNC-CH, UNC-C, NCSU and NCAT. We see the key to this effort stemming from PreMiEr advancing a bold and reimagined research agenda that not only benefits the historically privileged but also addresses the needs of historically marginalized communities. We recognize that while diversity is known to impact creativity by fueling innovation, diversity without an inclusive culture and equitable framework can significantly limit creativity and decrease participants’ commitment levels. Thus, our primary goal is to develop an inclusive and equitable culture where voices from diverse stakeholders are acknowledged and all participants are treated with respect and their ideas evaluated with the highest standards of scientific integrity.
PreMiEr’s commitment to DCI is evidenced by the composition of our ERC. Our leadership team is composed of 60% females and 20% URM faculty, while our senior personnel is composed of 43.6% females and 12.8% faculty from racially marginalized communities. We are committed to promoting an equitable academic and social environment where minority scholars can flourish. This will be achieved through leadership development for ERC leaders/mentors including effective communication strategies, fostering a thriving & engaging environment, and promoting an inclusive environment; and 3) feedback from PreMiEr’s advisory boards to our Executive and Steering Committees.
The four Diversity and Culture of Inclusion goals are to:
- Continue to prioritize and increase diversity, equity and inclusion across PreMiEr’s institutional ecosystem
- Promote an equitable academic and social environment where minority scholars can flourish
- Prioritize the recruitment and retention of diverse trainees
- Broaden participation in microbiome sciences
Award Abstract # 2133504: NSF Engineering Research Center for Precision Microbiome Engineering (PreMiEr)
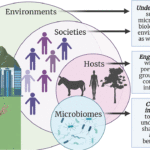
Microbes have colonized and adapted to most every environment on Earth, including the built environments that humans have created, such as the homes where we live and the pipes that bring us drinking water. It has been well established that microbial communities, or microbiomes, that colonize people have a direct influence on human health. The microbiome of the built environment, in particular, has gained increasing recognition for its key role in human health through its interaction with the human microbiome. However, despite this knowledge, no systematic infrastructure exists to decipher how microbial systems adapt to and grow within built environments, impeding our ability to diagnose built environment health and harness the power inherent to those microbiomes.
The Engineering Research Center for Precision Microbiome Engineering (PreMiEr) will create microbiome-based diagnostic tools and develop microbiome engineering approaches to monitor and operate built environments that maximize human health protection. Informed by societal needs and research-stakeholder teams, PreMiEr’s research design will work to prevent the spread of infectious agents, promote the colonization of beneficial microorganisms, and lead to strategies for controlling pandemics and antibiotic resistance—phenomena that have led to over six million deaths worldwide (as of June 2022) and cost the global economy an estimated $8 trillion in the last year alone. Integral to its research vision, PreMiEr will create diverse and inclusive interdisciplinary research and training hubs where engineers, microbiologists, social scientists, and ethicists work alongside theorists, model builders, and computational scientists to develop technologies that enable transformative engineering discoveries in safe, sustainable and responsible ways.
Our capacity to engineer microbiomes requires a fundamental understanding of concepts of community ecology and an ability to track, control, and model those interactions. To apply microbiome engineering to real-world systems, community level interactions must be integrated into a comprehensive, scalable modeling framework that requires iterative evaluation and validation in model testbeds. PreMiEr’s research organization is designed to generate fundamental understanding across these levels and functionalities, culminating in the development of a framework that enables the biodesign of smart and healthy built environments. PreMiEr will leverage advances in high-throughput genomic sequencing, high-resolution mass spectrometry, computational performance, and statistical modeling to unravel previously unknown mechanistic interactions. Enabling technologies will be developed to detect and define interactions in the built environment, including approaches that probe microbial dark matter for the development of built-environment health diagnostic tools; methods for targeted delivery of desired genetic features and microbial vectors; tools for fine in situ functional tuning; and predictive scalable statistical microbiome engineering models that consider high dimensionality, sparsity, and heterogeneity. These new technology elements will enable us to test hypotheses related to microbiome assembly and function.
Importantly, by incorporating social scientists and ethicists into PreMiEr’s research framework, non-social scientists’ work will be informed by consideration of the ethical, societal, and policy implications of their microbiome engineering discoveries. Through rigorous evaluation and iterative refinement of curricula, and institutional practices designed to support a culture of convergence and the dissemination of findings, PreMiEr will contribute to best practices in domestic training. The PreMiEr ERC will include targeted recruitment of trainees from underrepresented groups, building upon existing partnerships with our nation’s largest HBCU, and will provide immersion in research and training at the interface of multiple disciplines to address complex challenges. PreMiEr will train the next generation of diverse and highly motivated engineers and scientists in technical and professional skills to compete in the emerging arenas of microbial science and engineering. Ultimately, our work will advance collaborations and discovery focused on environmental microbiomes to engineer healthy built environments.
NC State PreMiEr Faculty

With backgrounds ranging from biomolecular engineering to fungal microbiomes, researchers from five NC State colleges are contributing their expertise to the National Science Foundation Engineering Research Center for Precision Microbiome Engineering.
| Photo | Faculty |
|---|---|
 | Jennifer KuzmaDr. Kuzma is the Goodnight-NCGSK Foundation Distinguished Professor in the School of Public and International Affairs in the College of Humanities and Social Sciences, Co-Director of the Genetic Engineering and Society Center, and a member of the Chancellor's Faculty Excellence Program
|
 | Yi-Hui ZhouDr. Zhou is an Associate Professor of Biological Sciences in the College of Sciences, Associate Member of the Department of Statistics, Associate Editor of Biostatistics, Associate Director of Outreach with the Bioinformatics Research Center, and a member of the Chancellor's Faculty Excellence Program
|
 | Benjamin CallahanDr. Callahan is an Assistant Professor of Microbiomes and Complex Microbial Communities in the Population Health and Pathobiology department of the College of Veterinary Medicine, a member of the Chancellor's Faculty Excellence Program, and is also affiliated with the Bioinformatics Research Center
|
 | Nathan CrookDr. Crook is an Assistant Professor of Chemical and Biomolecular Engineering in the College of Engineering, PI of the Crook Lab
|
 | Kevin GarciaDr. Garcia is an Assistant Professor in the Department of Crop and Soil Sciences in the College of Agriculture and Life Sciences and PI of the Garcia Lab
|
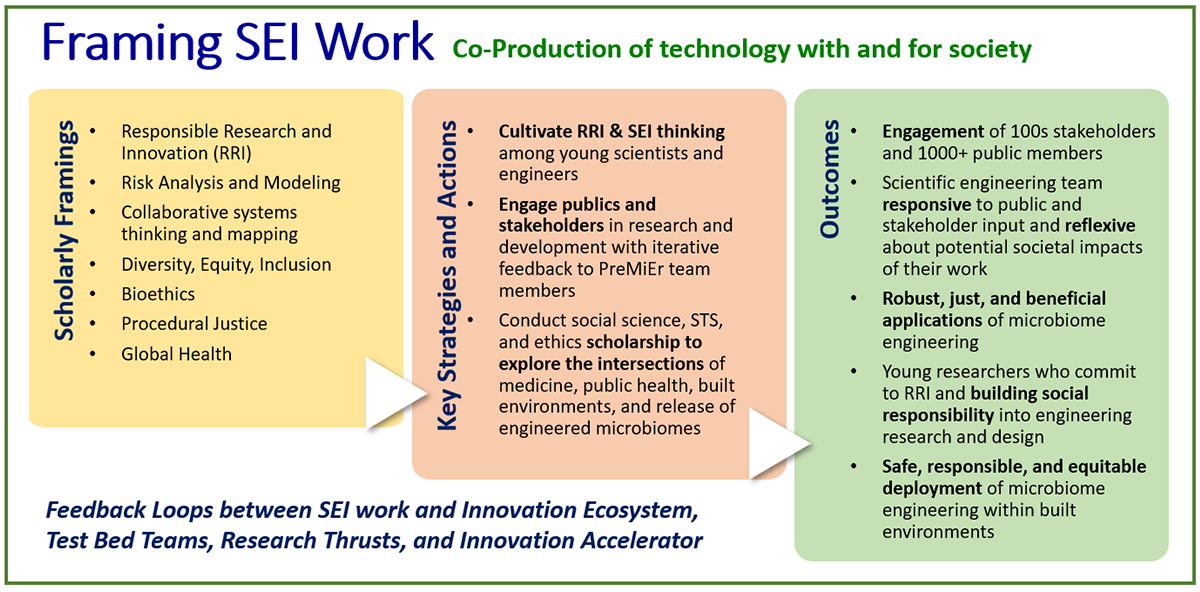
Social & Ethical Implications Team
PreMiEr’s third research core, focused on the social and ethical implications (SEI) of engineered microbiomes, evokes a diverse range of issues at the intersection of health and environmental risk, medical ethics, research ethics, environmental release of genetically modified organisms, public trust and perceptions, social equity, gender and racial inequities, privacy and regulation, and responsible governance.Woven into all of PreMiEr’s research activities, this provides a unique opportunity to engage researchers, engineers, stakeholders, and publics in emerging conversations about engineered microbiomes in built environments. It will also enable novel and ground-breaking scholarly examination of the various SEI aspects of PreMiEr’s research activities.

SEI Journal Club
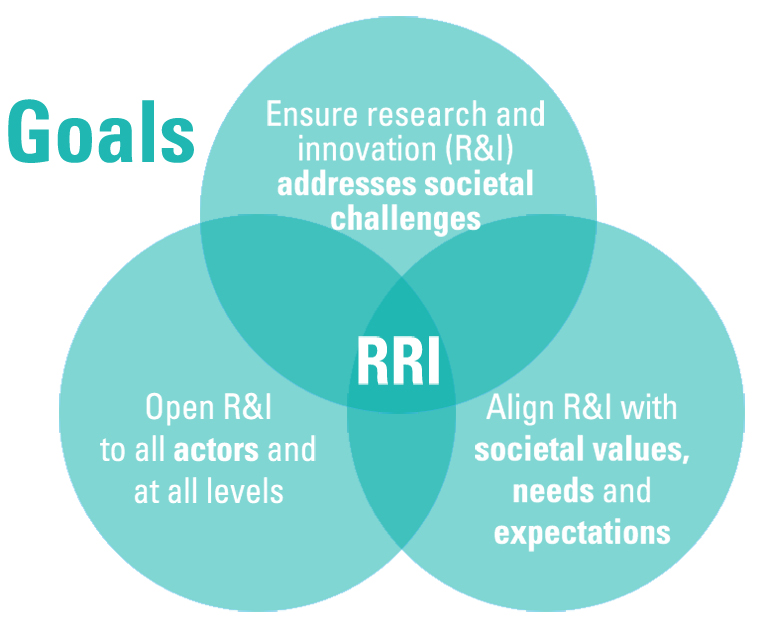
Responsible Research and Innovation
An interactive process by which societal actors and innovators become mutually responsive to each other with a view on the (ethical) acceptability, sustainability and societal desirability of the innovation process and its marketable products. (von Schomberg 2011)
SEI Action Plan & Projects
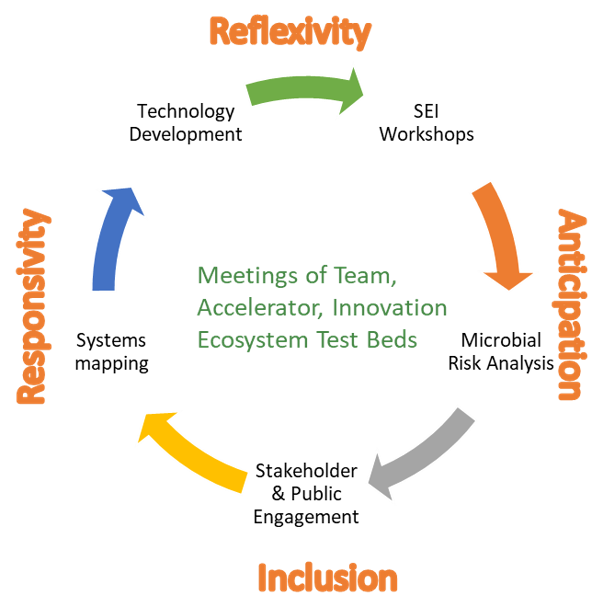
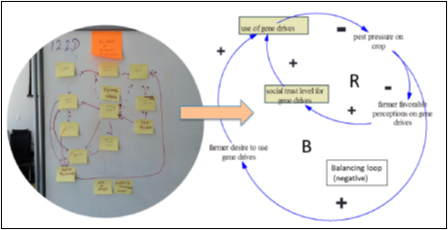
1) Collaborative Systems Mapping and Modeling—Convergence of Disciplines Across Team (Y1-Y5)
- Societal aspects, market barriers, microbial risk analysis, governance
- Integrated with Team Meetings of Innovation Accelerator, Test-Beds, and Innovation Ecosystem
- Part of Student Training and Short Courses; REU (research experience for undergraduates) Projects; and SEI Annual Workshops
- Provides Mechanisms and Framework for Iterative Feedback from Stakeholder and Public Engagement Projects to Research Thrusts, Cores and Innovation Accelerator
2) Public Engagement for Inclusion of Diverse Voices
- Draw on engagement infrastructure and experience of Core C partner centers
- Deliberative workshops on SEI and educational demonstrations at community labs across U.S. (Y3-5)
- Public dialogues and interviews of participants in and near Test Beds (Y2-5)
- Specific inclusion of perspectives from underserved racial groups
- Targeted inclusion of research participants (e.g. those with wearable devices)
- Feedback to research and engineering team via Innovation Accelerator, Test Beds, Innovation Ecosystem, Research Thrusts, and other team meetings
3) SEI Research and Deliberative Workshops
- Engage SEI experts around U.S. and world (Y1-5)
- National SEI Conferences (Y3 & 5)
- Engage junior SEI scholars, natural scientists and engineers
- Provide a network of professional development in RRI
- Special edition of journals and policy forum outputs
- SEI Expert Workshops (Y2 & 4)
- Bring in SEI expertise in addition to Core C leadership
- Additional risk analysts, legal scholars, economists, etc.
- Focused Risk Assessment Track in each workshop
- Become “premier” place for SEI scholarship and practice for microbiome engineering in built environments
- Be a national policy voice for built environments and microbiome engineering
4) Assessing Public and Stakeholder Attitudes
- Annual quantitative surveys with nationally representative group
- In-depth interviews with stakeholders on innovation ecosystem, market forces, and regulation & governance
- In-depth interviews with people in test bed areas on hopes, concerns, privacy and informed consent
- Focus groups and deliberative events at community labs
- Particular assessment of attitudes of historically marginalized groups
- Feedback to research team and industry stakeholders in Innovation Accelerator, Test Beds, Research Thrusts and Innovation Ecosystem Core
Key Outcome
Enhance the success of microbiome technology within society and its integration in society in responsible, just, and inclusive ways.
PreMiEr News
NC State part of $26 million grant to study microbiomes
Heidi Reid, September 7, 2022 | NC State is taking part in the National Science Foundation Engineering Research Center for Precision Microbiome Engineering (PreMiEr) to research genetically engineered microbiomes.
Exploring the Social, Ethical Sides of Microbiome Engineering
Nash Dunn, September 7, 2022 | At NSF center, NC State to Lead Research on Societal and Ethical Implications of Emerging Technologies
NC State to Research Implications of Engineered Microbiomes with New NSF Center Grant
Deborah Strange, August 10, 2022 | NC State University is part of a five-year, $26 million National Science Foundation center researching microbiome engineering.